understandings
● A gene is a heritable factor that consists of a length of DNA and influences a specific characteristic. ● A gene occupies a specific position on a chromosome.
● The various specific forms of a gene are alleles.
● Alleles differ from each other by one or only a few bases.
● New alleles are formed by mutation.
● The genome is the whole of the genetic information of an organism.
● The entire base sequence of human genes was sequenced in the Human Genome Project
● The various specific forms of a gene are alleles.
● Alleles differ from each other by one or only a few bases.
● New alleles are formed by mutation.
● The genome is the whole of the genetic information of an organism.
● The entire base sequence of human genes was sequenced in the Human Genome Project
applications and skills
- Application: The causes of sickle cell anaemia, including a base substitution mutation, a change to the base sequence of mRNA transcribed from it, and a change to the sequence of a polypeptide in haemoglobin.
- Application: Comparison of the number of genes in humans with other species.
- Skill: Use of a database to determine differences in the base sequence of a gene in two species
guidance
- Students should be able to recall one specific base substitution that causes glutamic acid to be substituted by valine as the sixth amino acid in the haemoglobin polypeptide.
- The number of genes in a species should not be referred to as genome size as this term is used for the total amount of DNA. At least one plant and one bacterium should be included in the comparison, and at least one species with more genes and one with fewer genes than a human.
- The GenBank® database can be used to search for DNA base sequences. The cytochrome c gene sequence is available for many different organisms and is of particular interest because of its use in reclassifying organisms into three domains.
- Deletions, insertions, and frame shift mutations do not need to be included.
what is a gene?
A gene is a heritable factor that consists of a length of DNA and influences a specific characteristic. ‘Heritable’ means passed on from parent to offspring, and ‘characteristic’ refers to genetic traits such as your hair colour or your blood type. The estimated 21 000 genes that you possess are organized into chromosomes.
A gene is found at a particular locus on a chromosome
A gene is found at a particular locus on a chromosome
- A gene for a specific trait occupies a corresponding place, called a locus (plural loci), on a chromosome.
- When geneticists map out the sequences of DNA, they carefully map the locus of each sequence. When further research reveals that a particular sequence controls a certain heritable factor, the locus of the gene is noted for further reference.
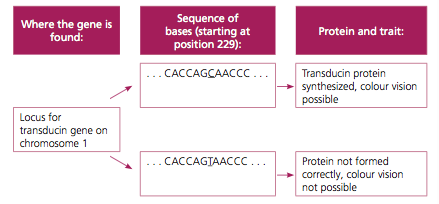
- For example, scientists now know that the locus of the gene controlling a protein called transducin that enables colour vision is found on chromosome 1.
- A mutation of this gene stops a person from being able to make the protein transducin properly, which is necessary to transmit information about colour from the eye to the brain; as a result, the person will not see in colour.
- This is an extremely rare genetic condition called complete a chromatopsia. When we say ‘the ability to see in colour is a genetic trait’ we mean one of two things is happening with someone’s DNA: either that person has the DNA code for making colour vision possible or the person does not have it.
- You will recall that you possess two copies of each gene in your body: one copy from your mother and one from your father.
- As a result, if you could look at the locus of the transducin gene on one of the two copies of your first chromosome, for example, you would find the same gene at the same locus on the other copy of chromosome 1.
- One copy would be the one your mother gave you and the other would be the copy your father gave you. Would those genes be identical? Not necessarily, because genes can come in different forms.
Alleles: versions of genes
- Variations or versions of a gene are called alleles.
- An allele is one specific form of a gene, differing from other alleles by one or a few bases.
- In the example of transducin and colour vision above, a single base pair difference between the most common allele (with a C at position 235) and the rare mutated allele (with a T at position 235) is all that is takes to determine whether you can distinguish colours or not.
- These different forms allow for a single trait, such as the trait for the ability to see in colour, to have variants, in this example either colour or grey-scale vision. Another example of the difference between two alleles of the same trait is the difference that causes the genetic condition cystic fibrosis.
Cystic FIBROSIS
- Maintaining a proper balance of fluids in the body is essential for good health.
- One such fluid is mucus, a thick, slippery, substance used in many parts of the body, including the lungs and intestines.
- A gene called CFTR, found on chromosome 7, plays a key role in the production of mucus.
- The standard version of this gene (the standard allele) allows a person’s mucus-producing cells to function properly, whereas an allele generated by a mutation of the CFTR gene causes cystic fibrosis.
- People with this genetic condition produce abnormally excessive quantities of mucus in various organs and have difficulties with their respiratory and digestive systems, among other complications.
- In this example, the trait is for mucus production; one allele is for a balanced mucus production, the other for excessive mucus production that leads to cystic fibrosis.
- We will see later how to calculate the chances of a child inheriting this condition from his or her parents.
One base can make a big difference
- From the sections on transcription and translation of DNA, you will remember how important it is for each letter in the genetic code to be in a specific place.
- If, for whatever reason, one or more of the bases (A, C, G or T) is misplaced or substituted for a different base, the results can be dramatic.
- As we have seen with cystic fibrosis, the difference between one version of a gene and another (the mutated and non-mutated alleles of the CFTR gene) can mean the difference between healthy organs and organs hampered by an overproduction of mucus.
- Another example of a change of bases can be seen in the gene ABCC11, which determines several things, one of them being whether or not the cerumen (ear wax) that you produce is wet or dry.
- Some people produce dry cerumen, which is flaky and crumbly with a grey colour, while others produce earwax that is more fluid and has an amber colour.
- The gene that determines this is on chromosome 16 and has two alleles: the G variant codes for dry cerumen, the A variant codes for wet cerumen.
- The allele containing G for wet earwax is much more common in European and African populations, while the allele containing A is much more common among Asians.
- Why is this of interest to geneticists? For one thing, it can reveal a lot about how populations have migrated and interbred in the past, but it can also reveal other things about our health.
- As curious as it may seem, the ABCC11 gene is also partly responsible for the smell of underarm sweat, as well as the production of breast milk, and could potentially have a link to breast cancer.
- Most women probably would not care whether or not they have the gene for dry or fluid earwax, but if they could find out whether they had an allele that could reduce their chances of having breast cancer, they might be much more interested.
mutations
A mutation is a random, rare change in genetic material. One type involves a change of the sequence of bases in DNA. If DNA replication works correctly, this should not happen . But nature sometimes makes mistakes. For example, the base thymine (T) might be put in the place of adenine (A) along the DNA sequence. When this happens, the corresponding bases along the messenger RNA (mRNA) are altered during transcription. As we have seen with the example of cystic fibrosis, mutated genes can have a negative effect on a person’s health. Sometimes, however, mutations can have a positive effect that is beneficial to an organism’s survival.
Are mutations good or bad for us?
LRP5 is a gene that helps immune system cells make a certain type of protein that acts as a receptor on their surfaces. Research indicates that this receptor is used by the human immunodeficiency virus (HIV) to infect the cells (see Section 6.3 for
a description of HIV). People with a mutation of LRP5 cannot make this receptor protein on their immune system’s cells and, as a result, HIV cannot infect them. This means that people with a mutated allele of LRP5 are naturally immune to HIV. Such a mutation is very rare in the human population.
A mutation that provides an individual or a species with a better chance for survival is considered to be a beneficial mutation, and there is a good chance that it will be passed on to the next generation. In contrast, mutations that cause disease or death are detrimental mutations, and they are less likely to be passed on to future generations, because they decrease the chances of an individual’s survival. In addition to beneficial and harmful mutations, there are neutral mutations that do not have an effect on a species’ survival.
When a mutation is successfully passed on from one generation to the next, it becomes a new allele: it is a new version of the original gene. This is how new alleles are produced. You and everyone you know possess many mutations. Whether they are harmful, beneficial or neutral depends on what they are and what kind of environment you need to survive in.
a description of HIV). People with a mutation of LRP5 cannot make this receptor protein on their immune system’s cells and, as a result, HIV cannot infect them. This means that people with a mutated allele of LRP5 are naturally immune to HIV. Such a mutation is very rare in the human population.
A mutation that provides an individual or a species with a better chance for survival is considered to be a beneficial mutation, and there is a good chance that it will be passed on to the next generation. In contrast, mutations that cause disease or death are detrimental mutations, and they are less likely to be passed on to future generations, because they decrease the chances of an individual’s survival. In addition to beneficial and harmful mutations, there are neutral mutations that do not have an effect on a species’ survival.
When a mutation is successfully passed on from one generation to the next, it becomes a new allele: it is a new version of the original gene. This is how new alleles are produced. You and everyone you know possess many mutations. Whether they are harmful, beneficial or neutral depends on what they are and what kind of environment you need to survive in.
A gene to help digestion
For most of our existence, humans have been hunter-gatherers and our genes are generally well adapted for this lifestyle. Originally, as for all mammals, the only age at which we drank milk was when we were infants. By the time our ancestors reached adulthood, their bodies had stopped being able to digest milk; more precisely, humans could not break down the disaccharide in milk called lactose. This continues to be the case for most of the human population today: more than half of the human population has lactose intolerance and those people can only digest lactose in their infancy. In the past 10 000 years, however, many human populations have adopted an agricultural- based lifestyle, raising animals for milk and consuming dairy products on a daily basis. In their genetic makeup, many agricultural societies show a higher frequency of the genetic code that allows humans to digest lactose throughout adulthood. From an evolutionary point of view, this advantage has increased humans’ ability to survive harsh climatic conditions. As European human populations spread out and established populations outside Europe, notably in North America, they brought their lactose tolerance (and their livestock) with them.
base substitution mutation
The type of mutation that results in a single letter being changed is called a base substitution mutation. The consequence of changing one base could mean that a different amino acid is placed in the growing polypeptide chain. This may have little or no effect on the organism, or it may have a major influence on the organism’s physical characteristics.
Sickle cell disease
In humans, a mutation is sometimes found in the gene that codes for haemoglobin in red blood cells. This mutation gives a different shape to the haemoglobin molecule. The difference leads to red blood cells that look very different from the usual flattened disc with a hollow in the middle.
The mutated red blood cell, with a characteristic curved shape, made its discoverers think of a sickle (a curved knife used to cut tall plants). The condition that results from this mutation is therefore called sickle cell disease, also known as sickle cell anaemia.
The kind of mutation that causes sickle cells is a base substitution mutation. If you look back at the two sequences given previously in the worked example on page 117, the first is for the section of the haemoglobin gene’s DNA that codes for standard-shaped red blood cells, whereas the second sequence shows the mutation that leads to the sickle shape. In this case, one base is substituted for another so that the sixth codon in this sequence of haemoglobin, GAG, becomes GTG. As a result, during translation, instead of adding glutamic acid, which is the intended amino acid in the sixth position of the sequence, valine is added there instead. Again, refer back to the worked example to see this mutation.
Because valine has a different shape and different properties compared with glutamic acid, the shape of the resulting polypeptide chain is modified. As a result of this, the haemoglobin molecule has different properties that cause the complications associated with sickle cell disease.
The symptoms of sickle cell disease are weakness, fatigue, and shortness of breath. Oxygen cannot be carried as efficiently by the irregularly shaped red blood cells. In addition, the haemoglobin tends to crystallize within the red blood cells, causing them to be less flexible. The affected red blood cells can get stuck in capillaries, so blood flow can be slowed or blocked, a condition that is painful for the sufferer.
People affected by sickle cell anaemia are at risk of passing the mutated gene on to their offspring. From a demographics point of view, the mutated gene is mostly found in populations originating from West Africa or from the Mediterranean.
A genome
How do we know all that we do about genes? How do we know where they are and what they do? Before answering these questions, it is important to appreciate the point that, although we have made considerable progress in the past few decades, our maps of human chromosomes are still far from complete, and there are many DNA sequences for which we do not know the function. As an analogy, think of the maps produced by cartographers and explorers in the Middle Ages; many parts of the globe remained uncharted and had the words terra incognita (Latin for ‘unknown land’) inscribed on them.
Sequencing DNA
In order to find out which gene does what, a list must be made showing the order of all the nucleotides in the DNA ode. Researchers use highly specialized laboratory equipment including sequencers to locate and identify sequences of bases. The complete set of an organism’s base sequences is called its genome.
A short fragment of a sequence looks like this: GTGGACCTGACTCCTGAGGAG. Each letter represents one of the four bases in the DNA code. This short fragment contains seven codons with a total of 21 bases represented by letters. Now imagine 3 billion of those letters: what would that look like? If you printed out 3000 base letters per page, it would need 1 million pages, which would stack about 100 m high. That’s an impressive quantity of information, especially considering that you can keep it all in the nucleus of a typical cell in your body.
The complete genomes of some organisms have been worked out. Among those organisms are the fruit fly, Drosophila melanogaster, and the bacterium, Escherichia coli, because these two organisms have been used extensively in genetics experiments for decades.
How do geneticists work out the complete genome?
Many steps are necessary. Here is a summary of one way of doing it: the Sanger technique.
Sickle cell disease
In humans, a mutation is sometimes found in the gene that codes for haemoglobin in red blood cells. This mutation gives a different shape to the haemoglobin molecule. The difference leads to red blood cells that look very different from the usual flattened disc with a hollow in the middle.
The mutated red blood cell, with a characteristic curved shape, made its discoverers think of a sickle (a curved knife used to cut tall plants). The condition that results from this mutation is therefore called sickle cell disease, also known as sickle cell anaemia.
The kind of mutation that causes sickle cells is a base substitution mutation. If you look back at the two sequences given previously in the worked example on page 117, the first is for the section of the haemoglobin gene’s DNA that codes for standard-shaped red blood cells, whereas the second sequence shows the mutation that leads to the sickle shape. In this case, one base is substituted for another so that the sixth codon in this sequence of haemoglobin, GAG, becomes GTG. As a result, during translation, instead of adding glutamic acid, which is the intended amino acid in the sixth position of the sequence, valine is added there instead. Again, refer back to the worked example to see this mutation.
Because valine has a different shape and different properties compared with glutamic acid, the shape of the resulting polypeptide chain is modified. As a result of this, the haemoglobin molecule has different properties that cause the complications associated with sickle cell disease.
The symptoms of sickle cell disease are weakness, fatigue, and shortness of breath. Oxygen cannot be carried as efficiently by the irregularly shaped red blood cells. In addition, the haemoglobin tends to crystallize within the red blood cells, causing them to be less flexible. The affected red blood cells can get stuck in capillaries, so blood flow can be slowed or blocked, a condition that is painful for the sufferer.
People affected by sickle cell anaemia are at risk of passing the mutated gene on to their offspring. From a demographics point of view, the mutated gene is mostly found in populations originating from West Africa or from the Mediterranean.
A genome
How do we know all that we do about genes? How do we know where they are and what they do? Before answering these questions, it is important to appreciate the point that, although we have made considerable progress in the past few decades, our maps of human chromosomes are still far from complete, and there are many DNA sequences for which we do not know the function. As an analogy, think of the maps produced by cartographers and explorers in the Middle Ages; many parts of the globe remained uncharted and had the words terra incognita (Latin for ‘unknown land’) inscribed on them.
Sequencing DNA
In order to find out which gene does what, a list must be made showing the order of all the nucleotides in the DNA ode. Researchers use highly specialized laboratory equipment including sequencers to locate and identify sequences of bases. The complete set of an organism’s base sequences is called its genome.
A short fragment of a sequence looks like this: GTGGACCTGACTCCTGAGGAG. Each letter represents one of the four bases in the DNA code. This short fragment contains seven codons with a total of 21 bases represented by letters. Now imagine 3 billion of those letters: what would that look like? If you printed out 3000 base letters per page, it would need 1 million pages, which would stack about 100 m high. That’s an impressive quantity of information, especially considering that you can keep it all in the nucleus of a typical cell in your body.
The complete genomes of some organisms have been worked out. Among those organisms are the fruit fly, Drosophila melanogaster, and the bacterium, Escherichia coli, because these two organisms have been used extensively in genetics experiments for decades.
How do geneticists work out the complete genome?
Many steps are necessary. Here is a summary of one way of doing it: the Sanger technique.
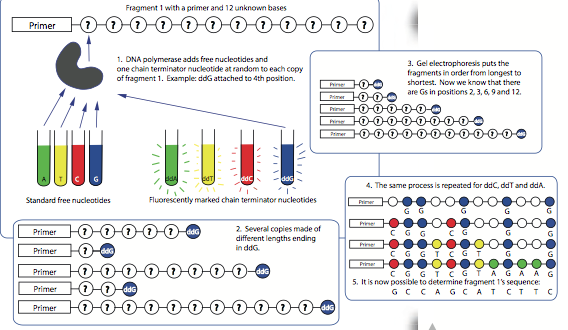
- Once a DNA sample has been taken, it is chopped up into fragments and copies are made of the fragments. A primer sequence is added to help start the process.
- To determine the sequence, a DNA polymerase enzyme attaches to one copy of the first fragment (let’s call it fragment 1). Then
it will start to add free nucleotides following the principle of complementary base pairing. Two kinds of nucleotides will be added.
- Some free nucleotides are standard ones and others are special dideoxynucleotide triphosphates (ddNTPs labelled ddA, ddT, ddC, and ddG in Figure 3.7) added as DNA chain terminators, meaning that when one is reached, the elongation of the strand is stopped. These have been previously marked with fluorescent markers to
identify them. Sometimes the chain termination happens all the way at the end of fragment 1, but most of the time the process stops before it reaches the end. This process happens on each of the many copies of fragment 1.
- The result is a series of new strands, some dozens of bases long, others only a handful of bases long, and some that have all the bases of fragment 1.
- Now everything is ready for the sequencing: the multiple chains of varying lengths (each with a fluorescently marked end) are placed in order from longest to shortest. This is done using a technique called gel electrophoresis, which will be explained in Section 3.5.
- To recognize each letter, a laser activates the fluorescent markers on the nucleotides as they go through the process. A sensor hooked up to a computer analyses the wavelength of the light and determines whether it represents an A, T, C, or G.
- The process must be repeated many times – for A, for T, C, and G. Repetitions make sure there are no errors. Fortunately, many copies of fragment 1 were made, so this is easy to check.
- Once fragment 1 is done, the lab technicians must process fragment 2, fragment 3, and so on, until all the fragments of the original sequence have been processed
At this stage, the challenge is to put all the sequenced fragments of code together. When the original sequence was chopped up to make all these fragments, they became mixed up and out of order. Now that we know what their sequences are,
we need to know the order in which to put them. This daunting task has been made easier by computers, but it consists of lining up any overlapping segments until they all match.
Since the Sanger technique was invented, many techniques have been developed to analyse each fragment only once, making it unnecessary to make multiple copies of each. This reduces the time and the cost of sequencing a genome. The objective of developing new sequencing techniques is to have a fast, inexpensive way to map anyone’s genome.
we need to know the order in which to put them. This daunting task has been made easier by computers, but it consists of lining up any overlapping segments until they all match.
Since the Sanger technique was invented, many techniques have been developed to analyse each fragment only once, making it unnecessary to make multiple copies of each. This reduces the time and the cost of sequencing a genome. The objective of developing new sequencing techniques is to have a fast, inexpensive way to map anyone’s genome.
The Human Genome Project
In 1990, an international cooperative venture called the Human Genome Project set out to sequence the complete human genome. Because the genome of an organism is a catalogue of all the bases it possesses, the Human Genome Project hoped to determine the order of all the bases A, T, C, and G in human DNA. In 2003, the Project announced that it had succeeded in achieving its goal. Now, scientists are working on deciphering which sequences represent genes and which genes do what. The human genome can be thought of as a map that can be used to show the locus of any gene on any one of the 23 pairs of chromosomes.
Before the Human Genome was mapped, fewer than 100 loci were known for genetic diseases. After the mapping was completed, more than 1400 were known, and today the number is in the thousands and increasing.
Before the Human Genome was mapped, fewer than 100 loci were known for genetic diseases. After the mapping was completed, more than 1400 were known, and today the number is in the thousands and increasing.
Using DNA to make medicines
Another advantageous use of the human genome is the production of new medications. This process involves several steps:
• find beneficial molecules that are produced naturally in healthy people
• find out which gene controls the synthesis of a desirable molecule
• copy that gene and use it as instructions to synthesize the molecule in a laboratory • distribute the beneficial therapeutic protein as a new medical treatment.
• find beneficial molecules that are produced naturally in healthy people
• find out which gene controls the synthesis of a desirable molecule
• copy that gene and use it as instructions to synthesize the molecule in a laboratory • distribute the beneficial therapeutic protein as a new medical treatment.
understandings
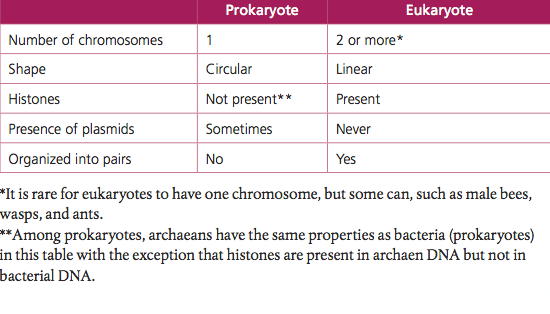
- Prokaryotes have one chromosome consisting of a circular DNA molecule.
- Some prokaryotes also have plasmids but eukaryotes do not.
- Eukaryote chromosomes are linear DNA molecules associated with histone proteins.
- In a eukaryote species there are different chromosomes that carry different gene
- Homologous chromosomes carry the same sequence of genes but not necessarily the same alleles of those genes.
- Diploid nuclei have pairs of homologous chromosomes.
- Haploid nuclei have one chromosome of each pair.
- The number of chromosomes is a characteristic feature of members of a species.
- A karyogram shows the chromosomes of an organism in homologous pairs of decreasing length.
- Sex is determined by sex chromosomes and autosomes are chromosomes that do not determine sex
applications and skills
Application: Cairns’ technique for measuring the length of DNA molecules by autoradiography.
● Application: Comparison of genome size in T2 phage, Escherichia coli, Drosophila melanogaster, Homo sapiens, and Paris japonica.
● Application: Comparison of diploid chromosome numbers of Homo sapiens, Pan troglodytes, Canis familiaris, Oryza sativa, and Parascaris equorum.
● Application: Use of karyograms to deduce sex and diagnose Down syndrome in humans.
● Skill: Use of databases to identify the locus of a human gene and its polypeptide product
● Application: Comparison of genome size in T2 phage, Escherichia coli, Drosophila melanogaster, Homo sapiens, and Paris japonica.
● Application: Comparison of diploid chromosome numbers of Homo sapiens, Pan troglodytes, Canis familiaris, Oryza sativa, and Parascaris equorum.
● Application: Use of karyograms to deduce sex and diagnose Down syndrome in humans.
● Skill: Use of databases to identify the locus of a human gene and its polypeptide product
guidance
● The terms karyotype and karyogram have different meanings. Karyotype is a property of a cell: the number and type of chromosomes present in the nucleus, not a photograph or diagram of them.
● Genome size is the total length of DNA in an organism. The examples of genome and chromosome number have been selected to allow points of interest to be raised.
● The two DNA molecules formed by DNA replication prior to cell division are considered to be sister chromatids until the splitting of the centromere at the start of anaphase. After this, they are individual chromosomes.
● Genome size is the total length of DNA in an organism. The examples of genome and chromosome number have been selected to allow points of interest to be raised.
● The two DNA molecules formed by DNA replication prior to cell division are considered to be sister chromatids until the splitting of the centromere at the start of anaphase. After this, they are individual chromosomes.
The chromosome in prokaryotes
You will recall from Chapter 1 that the nucleoid region of a bacterial cell contains a single, long, continuous, circular thread of DNA. Therefore, this region is involved with cell control and reproduction.
Notice how the presence of a single circular chromosome is a very different situation from all the cells we looked at in Section 3.1, which always had chromosomes in pairs. Why is this? Prokaryotes can reproduce using binary fission (dividing), whereas organisms such as plants and animals more frequently use sexual reproduction (involving a male and a female). Any time two parents are involved, the offspring will have pairs of chromosomes rather than single chromosomes. Because prokaryotes have only one parent, they have only one chromosome.
Some prokaryotes also have plasmids but eukaryotes do not
Escherichia coli, like many prokaryotes (bacteria), have small loops of DNA that are extra copies of some of the genetic material of the organism. These loops are called plasmids. These small, circular, DNA molecules are not connected to the main bacterial chromosome. The plasmids replicate independently of the chromosomal DNA. Plasmid DNA is not required by the cell under normal conditions, but it may help the cell adapt to unusual circumstances. Plasmids can also be found in Archaea as well as in bacteria. As we will see later in Section 3.5, these loops can be used in genetic engineering. Genetic manipulation using plasmids is not possible in eukaryotes such as plants and animals, because they do not have plasmids. Other techniques must be used for genetically modified (GM) crops and animals, which we will discuss later
EUKARYOTE CHROMOSOMES
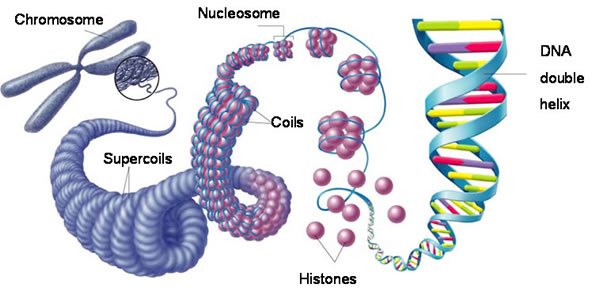
The DNA of eukaryotic cells most often occurs in the form of chromosomes. Chromosomes carry information necessary for the cell to exist. This allows the organism, whether unicellular or multicellular, to survive. DNA is the genetic material of the cell. It enables certain traits to be passed on to the next generation. When the cell is not dividing, the chromosomes are not visible structures. During this phase, the cell’s DNA is in the form of chromatin. Chromatin is formed of strands of DNA and proteins called histones.
multiple chromosomes
Homologous chromosomes: the same genes but not always the same alleles
- In a typical human cell, the 46 chromosomes can be grouped into 23 pairs of chromosomes called homologous chromosomes. Homologous means similar in shape and size, and it means that the two chromosomes carry the same genes. The example in Figure 3.9 shows one of the 23 pairs of homologous chromosomes found in humans.
- Remember that the reason there are two of each chromosome is that one came from the father and the other from the mother. Although a pair of homologous chromosomes carries the same genes, they are not identical because the alleles for the genes from each parent could be different. the locus shown contains different coloured bands, revealing that this individual got a different allele from his or her mother than from his or her father for this particular gene.
- Two chromosomes together as a single pair, but that each chromosome has been doubled as a result of DNA replication. Chromosomes only look like this when the cell they are in is getting ready to divide. At this stage, the two blue-banded zones are part of two connected sister chromatids forming a single chromosome attached at the centromere. Likewise, the two red-banded zones belong to two sister chromatids.
- Each chromatid includes the long arm as well as the short arm (the one that contains the coloured bands in this example). This will be important to remember later, when we watch the sister chromatids split during cell division. When the chromatids separate, they become two identical chromosomes. But as long as they are attached at the centromere, they are considered to be part of a single chromosome.
Sources
- Molecular Biology book chapter 3
- Powerpoint's shown in class